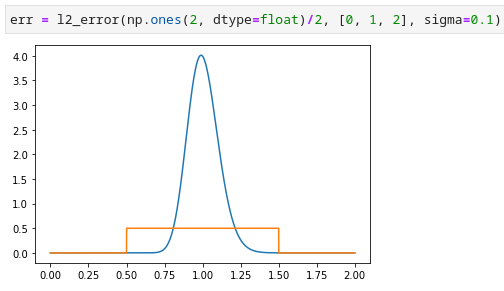
Here is a python script for calculating the L2 norm of a lognormal distribution.
from scipy.stats import lognorm
from scipy.interpolate import interp1d
import numpy as np
import matplotlib.pyplot as plt
TARGET_ERR = 1e-2
def l2_error(pmf: np.array, x_grid: np.array, sigma=0.1):
pmf = np.array(pmf)
x_grid = np.array(x_grid)
assert all(pmf >= 0)
assert np.isclose(sum(pmf), 1)
assert all(x_grid >= 0)
assert all(np.diff(x_grid) > 0)
assert len(pmf) + 1 == len(x_grid)
n_point = 2 ** 22
tail_value = (TARGET_ERR / 100) ** 2
min_x = lognorm.ppf(tail_value, sigma)
max_x = lognorm.ppf(1 - tail_value, sigma)
x_middle = np.linspace(min_x,max_x, n_point)
x_lower_tail = np.linspace(0, min_x, n_point//1000)
x_upper_tail = np.linspace(max_x, x_grid[-1], n_point//1000) if x_grid[-1] > max_x else np.array([])
x_approx = np.diff(x_grid) / 2 + x_grid[:-1]
x_approx = np.concatenate(([x_grid[0]], x_approx, [x_grid[-1]]))
pdf_approx = pmf /np.diff(x_grid)
pdf_approx = np.concatenate(([pdf_approx[0]], pdf_approx, [pdf_approx[-1]]))
fy = interp1d(x_approx, pdf_approx, kind='nearest', assume_sorted=True, fill_value=(0, 0), bounds_error=False)
x_full = np.concatenate((x_lower_tail[:-1], x_middle, x_upper_tail[1:]))
approx_pdf = fy(x_full)
full_pdf = lognorm.pdf(x_full, sigma)
dx = np.diff(x_full)
dx = np.append(dx, 0)
plt.plot(full_pdf)
plt.plot(approx_pdf)
upper_tail_err_2_approx = lognorm.sf(x_full[-1], sigma)
main_err_2 = sum((full_pdf - approx_pdf) ** 2 * dx)
err = (upper_tail_err_2_approx + main_err_2) ** 0.5
return err
def vanilla():
s = 0.1
qubits = 12
partial = 2 ** qubits
tail_value = (TARGET_ERR / 100) ** 2
x_approx = np.linspace(lognorm.ppf(tail_value, s),
lognorm.ppf(1 - tail_value, s), partial + 1)
pmf = np.diff(lognorm.cdf(x_approx, s))
pmf = pmf / sum(pmf)
print(l2_error(pmf, x_approx, sigma=s))
p = [0,
0,
0,
0,
0,
0.081,
0.02,
0.1,
0.5,
0.115,
0.15,
0.03,
0.004,
0,
0,
0]
x = [0.1,
0.2,
0.3,
0.4,
0.5,
0.6,
0.7,
0.8,
0.9,
1,
1.1,
1.2,
1.3,
1.4,
1.5,
1.6,
1.7]